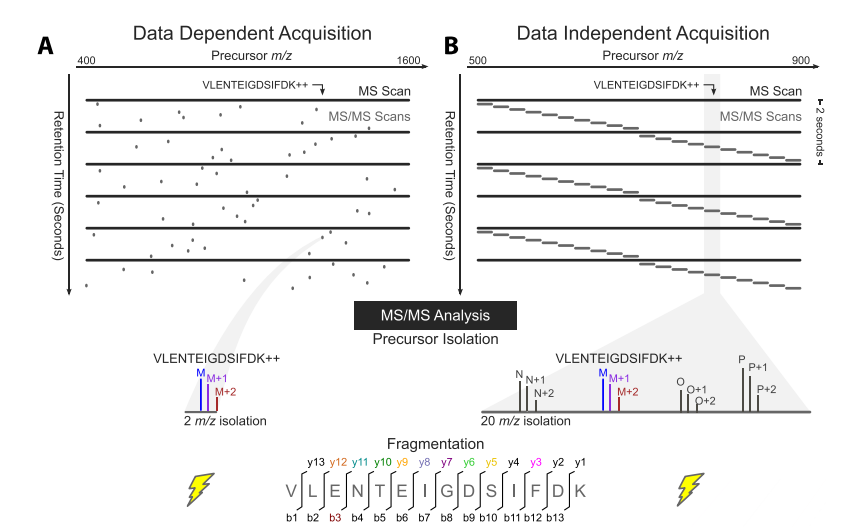
Data Dependent Acquisition and Data Independent Acquisition Mass Spectrometry - Creative Proteomics Blog

Figure 5 from A microelectromechanical systems-enabled, miniature triple quadrupole mass spectrometer. | Semantic Scholar

Comparison of linear intrascan and interscan dynamic ranges of Orbitrap and ion‐mobility time‐of‐flight mass spectrometers - Kaufmann - 2017 - Rapid Communications in Mass Spectrometry - Wiley Online Library

The BoxCar acquisition method a, The high dynamic range of the incoming... | Download Scientific Diagram
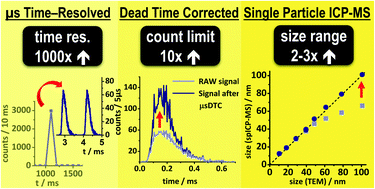
Single particle inductively coupled plasma mass spectrometry: investigating nonlinear response observed in pulse counting mode and extending the linear dynamic range by compensating for dead time related count losses on a microsecond

Comparison of linear intrascan and interscan dynamic ranges of Orbitrap and ion‐mobility time‐of‐flight mass spectrometers - Kaufmann - 2017 - Rapid Communications in Mass Spectrometry - Wiley Online Library

Improved Dynamic Range, Resolving Power, and Sensitivity Achievable with FT-ICR Mass Spectrometry at 21 T Reveals the Hidden Complexity of Natural Organic Matter | Analytical Chemistry
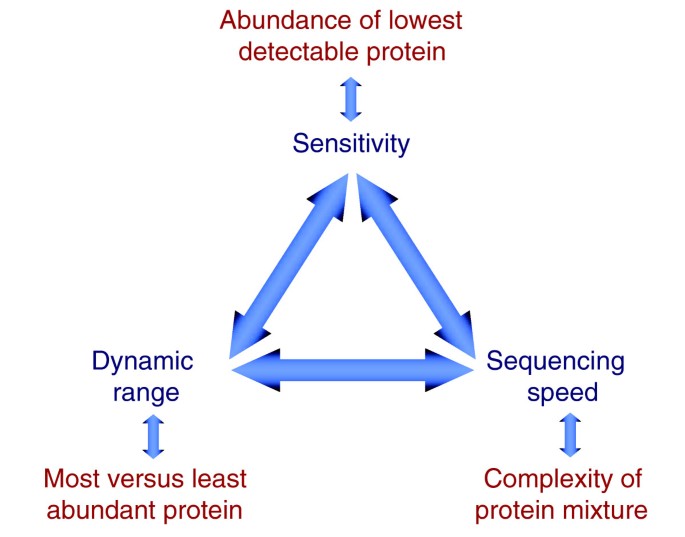
Status of complete proteome analysis by mass spectrometry: SILAC labeled yeast as a model system | Genome Biology | Full Text

Protein proportions and dynamic range in murine skeletal muscle tissue.... | Download Scientific Diagram

Expanding the linear dynamic range for quantitative liquid chromatography-high resolution mass spectrometry utilizing natural isotopologue signals - ScienceDirect

Single Particle Inductively Coupled Plasma Mass Spectrometry: Investigating Nonlinear Response Observed in Pulse Counting Mode and Extending the Linear Dynamic Range by Compensating for Dead Time Related Count Losses on a Microsecond

Comparison of linear intrascan and interscan dynamic ranges of Orbitrap and ion‐mobility time‐of‐flight mass spectrometers - Kaufmann - 2017 - Rapid Communications in Mass Spectrometry - Wiley Online Library

Improved Quantitative Dynamic Range of Time-of-Flight Mass Spectrometry by Simultaneously Waveform-Averaging and Ion-Counting Data Acquisition | Journal of The American Society for Mass Spectrometry

Figure 2 from Mass Spectrometry-Based Identification of Muscle-Associated and Muscle-Derived Proteomic Biomarkers of Dystrophinopathies. | Semantic Scholar
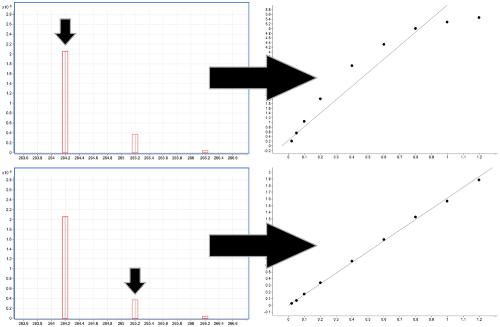
Increasing the linear dynamic range in LC–MS: is it valid to use a less abundant isotopologue?,Drug Testing and Analysis - X-MOL